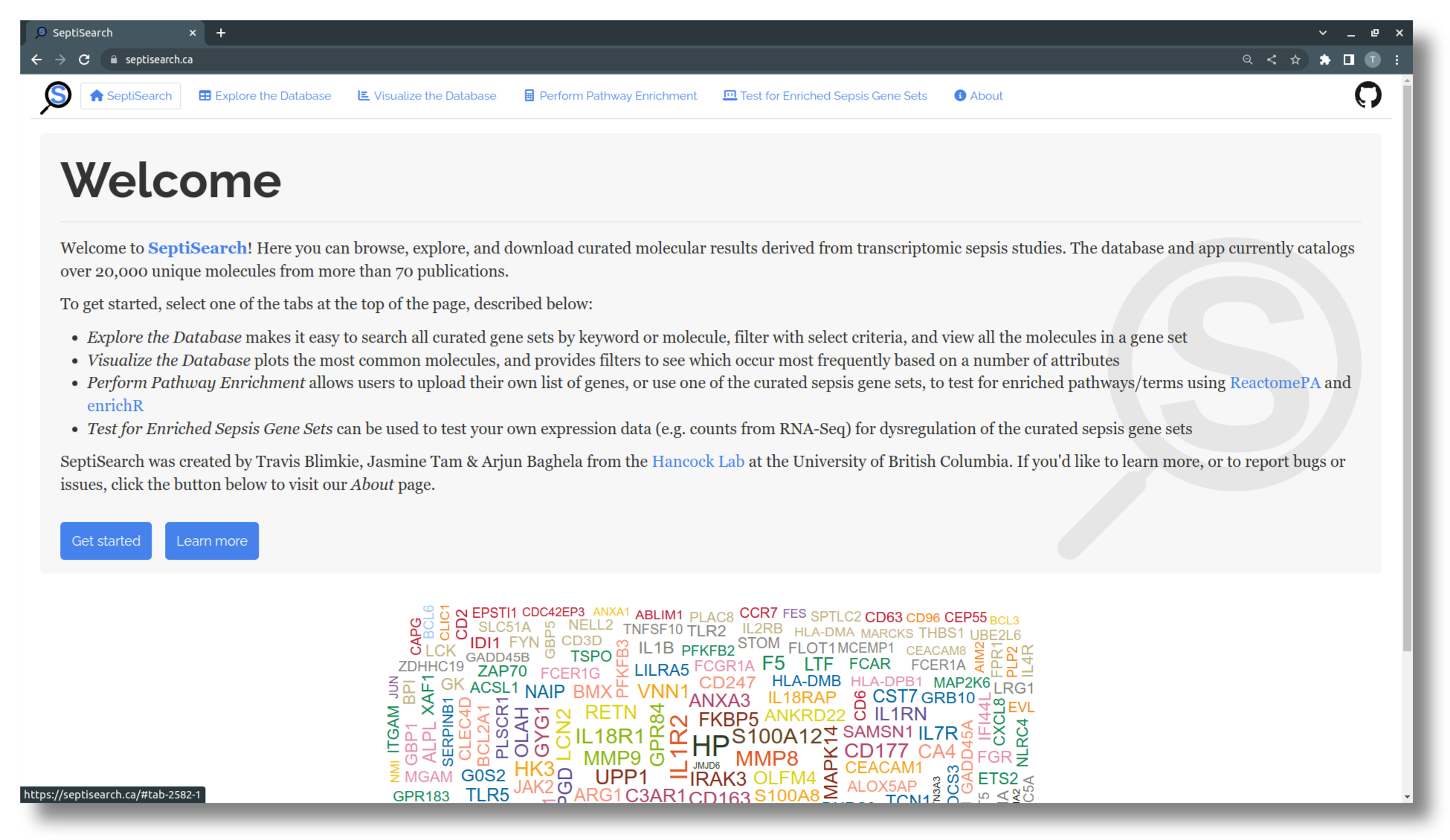
Welcome to the SeptiSearch tutorial pages
SeptiSearch is an interactive Shiny app providing access to manually-curated molecular sepsis data from current publications.
Visit septisearch.ca Go to the GitHub page
The sidebar on the left contains a section for each tab in SeptiSearch, plus a General Troubleshooting page for non-specific questions or issues.